Inference for multiple linear regression
February 05, 2026
Announcements
- Lab 02 due TODAY at 11:59pm
- HW 02 due Tuesday, February 10 at 11:59pm
- Lecture recordings + exam 01 practice problems on menu of course website
Topics
Interaction effects (from last class)
Inference for multiple linear regression
Computational setup
Data: Palmer penguins
The penguins data set contains data for penguins found on three islands in the Palmer Archipelago, Antarctica. Data were collected and made available by Dr. Kristen Gorman and the Palmer Station, Antarctica LTER, a member of the Long Term Ecological Research Network. These data can be found in the palmerpenguins R package.
# A tibble: 342 × 4
body_mass_g flipper_length_mm bill_length_mm species
<int> <int> <dbl> <fct>
1 3750 181 39.1 Adelie
2 3800 186 39.5 Adelie
3 3250 195 40.3 Adelie
4 3450 193 36.7 Adelie
5 3650 190 39.3 Adelie
6 3625 181 38.9 Adelie
7 4675 195 39.2 Adelie
8 3475 193 34.1 Adelie
9 4250 190 42 Adelie
10 3300 186 37.8 Adelie
# ℹ 332 more rowsVariables
Predictors:
bill_length_mm: Bill length in millimetersflipper_length_mm: Flipper length in millimetersspecies: Adelie, Gentoo, or Chinstrap species
Response: body_mass_g: Body mass in grams
The goal of this analysis is to use the bill length, flipper length, and species to predict body mass.
Interaction effects
Interaction terms
- Sometimes the relationship between a predictor variable and the response depends on the value of another predictor variable.
- This is an interaction effect.
- To account for this, we can include interaction terms in the model.
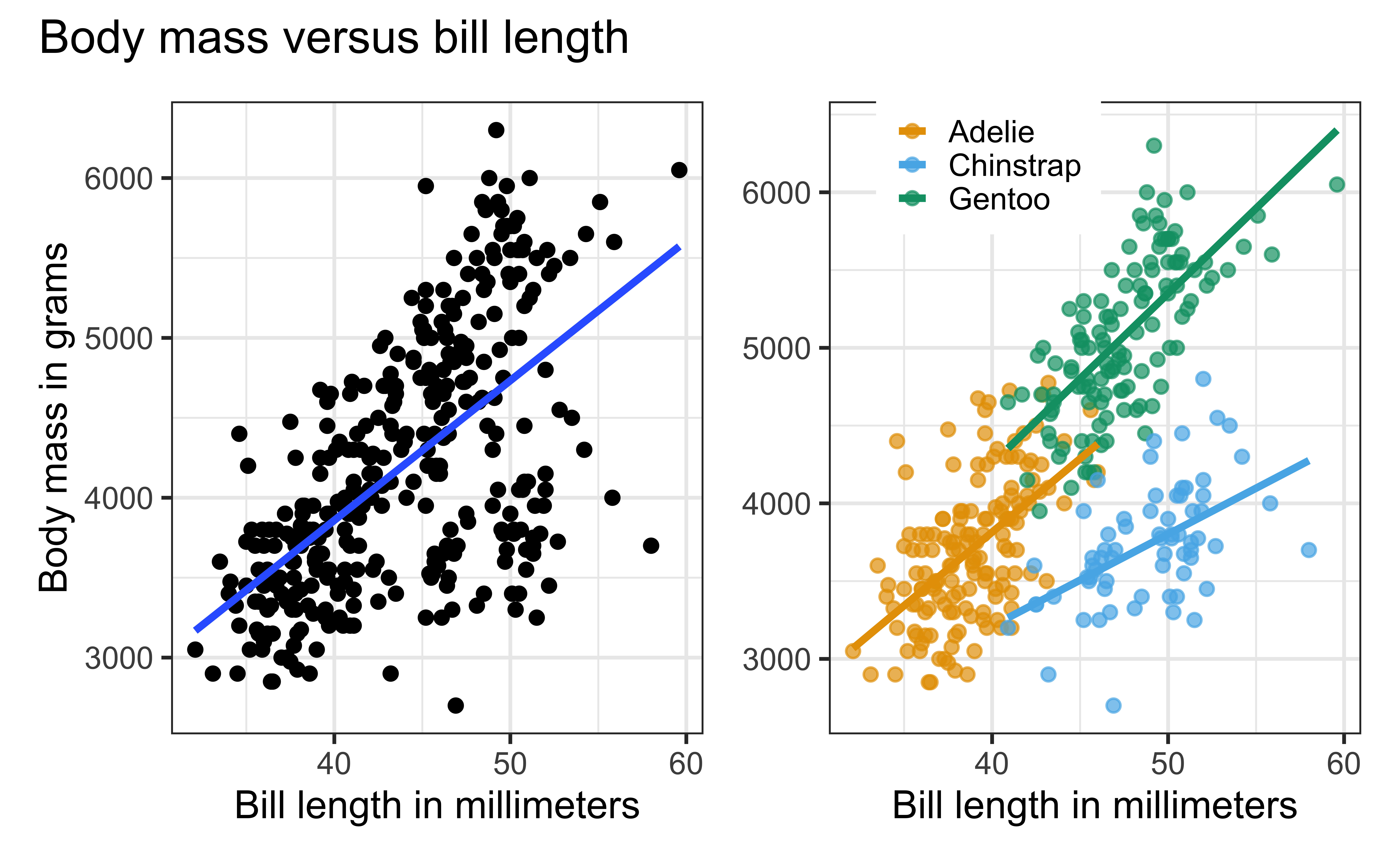
Bill length versus species
If the lines are not parallel, there is indication of a potential interaction effect, i.e., the slope of bill length may differ based on the species.
Interaction term in model
| term | estimate | std.error | statistic | p.value |
|---|---|---|---|---|
| (Intercept) | -4297.905 | 645.054 | -6.663 | 0.000 |
| flipper_length_mm | 27.263 | 3.175 | 8.586 | 0.000 |
| speciesChinstrap | 1146.287 | 726.217 | 1.578 | 0.115 |
| speciesGentoo | 54.716 | 619.934 | 0.088 | 0.930 |
| bill_length_mm | 72.692 | 10.642 | 6.831 | 0.000 |
| speciesChinstrap:bill_length_mm | -41.035 | 16.104 | -2.548 | 0.011 |
| speciesGentoo:bill_length_mm | -1.163 | 14.436 | -0.081 | 0.936 |
Interpreting interaction terms
- What the interaction means: The effect of bill length on the body mass is 41.035 less when the penguin is from the Chinstrap species compared to the effect for the Adelie species, holding all else constant.
- Interpreting
bill_length_mmfor Chinstrap: For each additional millimeter in bill length, we expect the body mass of Chinstrap penguins to increase by 31.657 grams (72.692 - 41.035), holding flipper length constant.
Inference for multiple linear regression
Inference framework
Our objective is to infer properties about a population using data from observational (or experimental) data collection
Pre data collection: Before collecting the data, the data are unknown and random. \(\hat{\beta}_j\), which is a function of the data, is also unknown and random.
Post data collection: After collecting the data, \(\hat{\beta}_j\) is fixed and known.
In all cases: The true population parameter, \(\beta_j\), is fixed but unknown.
Multiple linear regression
The multiple linear regression model assumes \[Y|X_1, X_2, \ldots, X_p \sim N(\beta_0 + \beta_1 X_1 + \beta_2 X_2 + \dots + \beta_p X_p, \sigma_\epsilon^2)\]
For a given observation \((x_{i1}, x_{i2}, \ldots, x_{ip}, y_i)\), we can rewrite the previous statement as
\[y_i = \beta_0 + \beta_1 x_{i1} + \beta_2 x_{i2} + \dots + \beta_p x_{ip} + \epsilon_{i} \hspace{10mm} \epsilon_i \sim N(0,\sigma_{\epsilon}^2)\]
Estimating \(\sigma_\epsilon\)
For a given observation \((x_{i1}, x_{i2}, \ldots,x_{ip}, y_i)\) the residual is \[e_i = y_{i} - (\hat{\beta}_0 + \hat{\beta}_1 x_{i1} + \hat{\beta}_{2} x_{i2} + \dots + \hat{\beta}_p x_{ip})\]
The estimated value of the regression standard error , \(\sigma_{\epsilon}\), is
\[\hat{\sigma}_\epsilon = \sqrt{\frac{\sum_{i=1}^ne_i^2}{n-p-1}}\]
As with SLR, we use \(\hat{\sigma}_{\epsilon}\) to compute \(SE_{\hat{\beta}_j}\), the standard error of the coefficient for predictor \(x_j\). See A.6: Distribution of model coefficients for more detail.
\(SE_{\hat{\beta}_j}\)
- \(SE_{\hat{\beta}_j} = \sqrt{Var_{\hat{\beta}_j}}\)
- Let’s look at \(Var_{\hat{\beta}_{flipper}, \hat{\beta}_{bill}}\) corresponding to the model \(\widehat{\text{body_mass_g}} = \hat{\beta}_0 + \hat{\beta}_1 \text{flipper_length_mm} + \hat{\beta}_2 \text{bill_length_mm}\)
(Intercept) flipper_length_mm bill_length_mm
(Intercept) 94838.825 -512.386 194.921
flipper_length_mm -512.386 4.045 -6.836
bill_length_mm 194.921 -6.836 26.831The diagonal elements are the variances for each coefficient
The off-diagonal elements are the covariances (measure of association) between the predictors
\(SE_{\hat{\beta}_j}\)
| term | estimate | std.error | statistic | p.value |
|---|---|---|---|---|
| (Intercept) | -5736.897 | 307.959 | -18.629 | 0.000 |
| flipper_length_mm | 48.145 | 2.011 | 23.939 | 0.000 |
| bill_length_mm | 6.047 | 5.180 | 1.168 | 0.244 |
Hypothesis test for \(\beta_j\)
Review: SLR hypothesis test
| term | estimate | std.error | statistic | p.value |
|---|---|---|---|---|
| (Intercept) | -5780.831 | 305.815 | -18.903 | 0 |
| flipper_length_mm | 49.686 | 1.518 | 32.722 | 0 |
- Set hypotheses: \(H_0: \beta_1 = 0\) vs. \(H_a: \beta_1 \ne 0\)
- Calculate test statistic and p-value: The test statistic is \(t= 32.722\). The p-value is calculated using a \(t\) distribution with 340 degrees of freedom. The p-value is \(\approx 0\) .
- State the conclusion: The p-value is small, so we reject \(H_0\). The data provide strong evidence that flipper length is a useful predictor of body mass, i.e. there is a linear relationship between the flipper length and body mass of penguins.
Multiple linear regression
| term | estimate | std.error | statistic | p.value |
|---|---|---|---|---|
| (Intercept) | -3904.387 | 529.257 | -7.377 | 0.000 |
| flipper_length_mm | 27.429 | 3.176 | 8.638 | 0.000 |
| speciesChinstrap | -748.562 | 81.534 | -9.181 | 0.000 |
| speciesGentoo | 90.435 | 88.647 | 1.020 | 0.308 |
| bill_length_mm | 61.736 | 7.126 | 8.664 | 0.000 |
MLR hypothesis test: flipper_length_mm
- Set hypotheses: \(H_0: \beta_{flipper} = 0\) vs. \(H_a: \beta_{flipper} \ne 0\), given species and bill length in the model
- Calculate test statistic and p-value: The test statistic is \(t = 8.638\). The p-value is computed from a \(t\) distribution with 337 df.
- State the conclusion: The p-value is small, so we reject \(H_0\). The data provide strong evidence that flipper length is a useful predictor of body mass, after accounting for bill length and species.
Interaction between flipper_length_mm and species?
| term | estimate | std.error | statistic | p.value |
|---|---|---|---|---|
| (Intercept) | -3341.616 | 810.145 | -4.125 | 0.000 |
| flipper_length_mm | 24.963 | 4.339 | 5.753 | 0.000 |
| speciesChinstrap | -27.293 | 1394.166 | -0.020 | 0.984 |
| speciesGentoo | -2215.913 | 1328.583 | -1.668 | 0.096 |
| bill_length_mm | 59.305 | 7.250 | 8.180 | 0.000 |
| flipper_length_mm:speciesChinstrap | -3.485 | 7.211 | -0.483 | 0.629 |
| flipper_length_mm:speciesGentoo | 11.026 | 6.482 | 1.701 | 0.090 |
Do the data provide evidence of a significant interaction effect? Comment on the significance of the interaction terms.
Confidence interval for \(\beta_j\)
Confidence interval for \(\beta_j\)
- The \(C\%\) confidence interval for \(\beta_j\) \[\hat{\beta}_j \pm t^* SE_{\hat{\beta}_j}\] where \(t^*\) follows a \(t\) distribution with \(n - p - 1\) degrees of freedom.
- Generically: We are \(C\%\) confident that the interval LB to UB contains the population coefficient of \(x_j\).
- In context: We are \(C\%\) confident that for every one unit increase in \(x_j\), \(y\) changes by LB to UB units, on average, holding all else constant.
Confidence interval for \(\beta_j\)
| term | estimate | std.error | statistic | p.value | conf.low | conf.high |
|---|---|---|---|---|---|---|
| (Intercept) | -3904.387 | 529.257 | -7.377 | 0.000 | -4945.450 | -2863.324 |
| flipper_length_mm | 27.429 | 3.176 | 8.638 | 0.000 | 21.182 | 33.675 |
| speciesChinstrap | -748.562 | 81.534 | -9.181 | 0.000 | -908.943 | -588.182 |
| speciesGentoo | 90.435 | 88.647 | 1.020 | 0.308 | -83.937 | 264.807 |
| bill_length_mm | 61.736 | 7.126 | 8.664 | 0.000 | 47.720 | 75.753 |
CI for flipper_length_mm
| term | estimate | std.error | statistic | p.value | conf.low | conf.high |
|---|---|---|---|---|---|---|
| (Intercept) | -3904.387 | 529.257 | -7.377 | 0.000 | -4945.450 | -2863.324 |
| flipper_length_mm | 27.429 | 3.176 | 8.638 | 0.000 | 21.182 | 33.675 |
| speciesChinstrap | -748.562 | 81.534 | -9.181 | 0.000 | -908.943 | -588.182 |
| speciesGentoo | 90.435 | 88.647 | 1.020 | 0.308 | -83.937 | 264.807 |
| bill_length_mm | 61.736 | 7.126 | 8.664 | 0.000 | 47.720 | 75.753 |
We are 95% confident that for every one millimeter increase in a penguin’s flipper length, its weight is greater by 21.182 to 33.675 grams, on average, holding bill length and species constant.
CI for speciesChinstrap
| term | estimate | std.error | statistic | p.value | conf.low | conf.high |
|---|---|---|---|---|---|---|
| (Intercept) | -3904.387 | 529.257 | -7.377 | 0.000 | -4945.450 | -2863.324 |
| flipper_length_mm | 27.429 | 3.176 | 8.638 | 0.000 | 21.182 | 33.675 |
| speciesChinstrap | -748.562 | 81.534 | -9.181 | 0.000 | -908.943 | -588.182 |
| speciesGentoo | 90.435 | 88.647 | 1.020 | 0.308 | -83.937 | 264.807 |
| bill_length_mm | 61.736 | 7.126 | 8.664 | 0.000 | 47.720 | 75.753 |
Interpret the 95% CI for
speciesChinstrap.Does this model provide evidence that the mean body mass of Chinstrap penguins differs from the mean body mass of Adelie penguins?
Inference pitfalls
Large sample sizes
Caution
If the sample size is large enough, the test will likely result in rejecting \(H_0: \beta_j = 0\) even \(x_j\) has a very small effect on \(y\).
Consider the practical significance of the result not just the statistical significance.
Use the confidence interval to draw conclusions instead of relying only p-values.
Small sample sizes
Caution
If the sample size is small, there may not be enough evidence to reject \(H_0: \beta_j=0\).
When you fail to reject the null hypothesis, DON’T immediately conclude that the variable has no association with the response.
There may be a linear association that is just not strong enough to detect given your data, or there may be a non-linear association.
Application exercise
Questions
Recap
- Introduced inference for multiple linear regression
Next class
- Model conditions and diagnostics
- Complete Lecture 10 prepare